Paper II
The Genetic Code from $\mathbb{Z}_8$
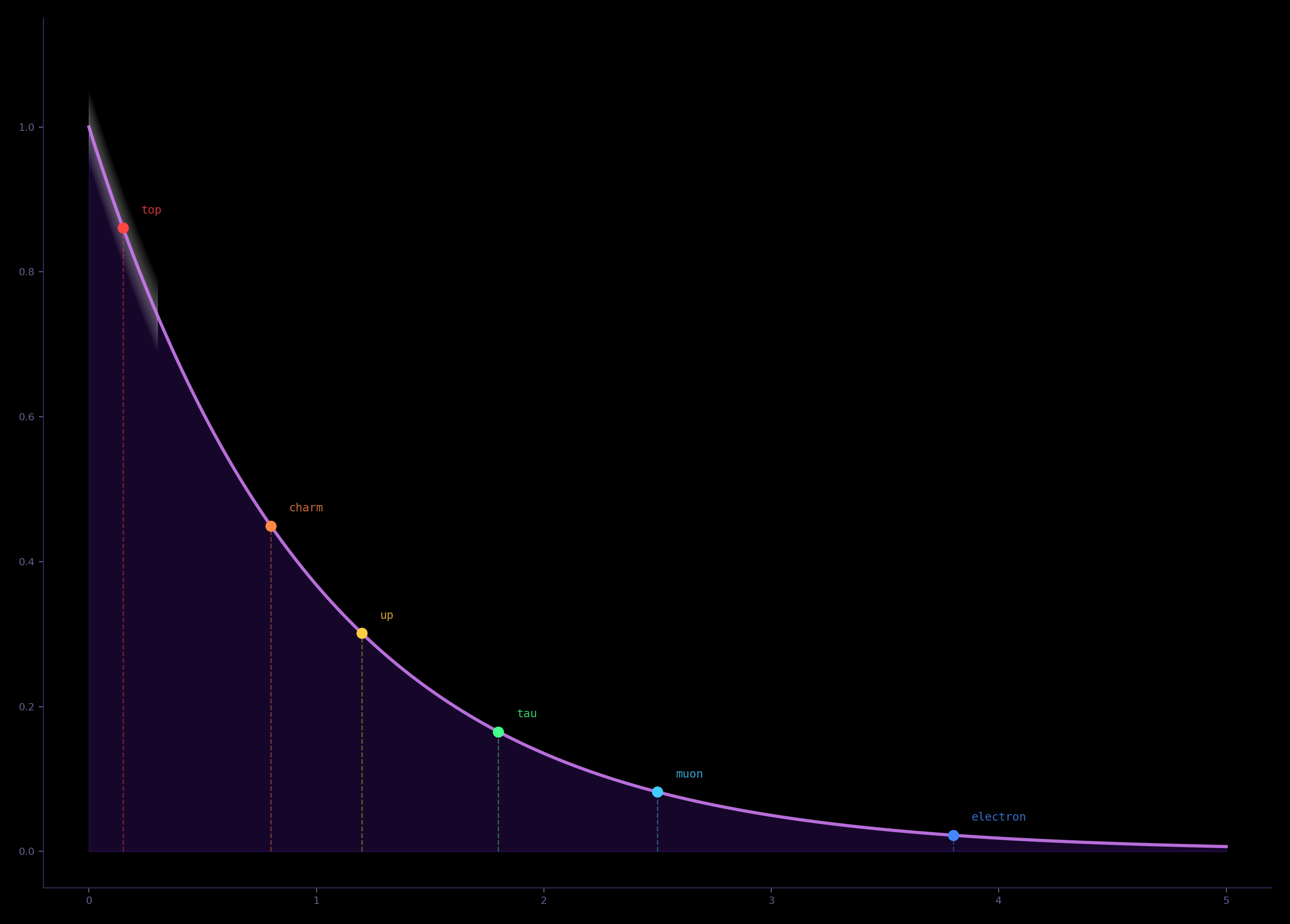
The $\ell = 0$ junction mode on $S^2 \!\vee\! S^2$ falls at 1.37 eV—the covalent bond energy
scale. The same $\mathbb{Z}_8$ holonomy that generates the particle spectrum reproduces the
combinatorial architecture of the genetic code. No additional parameters.
Combinatorial Structure
| $\mathbb{Z}_8$ Feature | Value | Genetic Code | Class |
|---|
| $\varphi(8)$ generators | 4 | DNA bases | D |
| Sectors at junction | 3 | Codon positions | D |
| Ordered triples $4^3$ | 64 | Total codons | D |
| $\binom{6}{3}$ multisets | 20 | Amino acids | D |
| $\text{Aut}(\mathbb{Z}_8)$ orbits | 5 | Degeneracy classes | D |
Watson–Crick: Two-Stage Filter
Stage 1 — Phase interference:
$|e^{i\phi_{j_1}} + e^{i\phi_{j_2}}|^2$ eliminates destructive pairs $(1,5)$ and $(3,7)$.
Stage 2 — Holonomy closure:
$j_1 + j_2 \equiv 0 \pmod{8}$ selects exactly $\{1,7\} = A\text{-}T$ and $\{3,5\} = G\text{-}C$.
| Pair | Interference | Closure | Classification |
|---|
| (1,7) = A-T | Constructive | Closed | Watson–Crick |
| (3,5) = G-C | Constructive | Closed | Watson–Crick |
| (1,3) | Constructive | Open | Mispair |
| (5,7) | Constructive | Open | Mispair |
| (1,5) | Destructive | Open | Forbidden |
| (3,7) | Destructive | Open | Forbidden |
Three Involutions: Klein Four-Group Saturation
$\mathbb{Z}_8^* \cong V_4 = \mathbb{Z}_2 \times \mathbb{Z}_2$ has exactly 3 non-trivial involutions.
Biology uses all three. No further involutions exist.
| Involution | Map | Orbits | Biological Function |
|---|
| $I_7$ | $j \to 7j$ | $\{1,7\},\{3,5\}$ | Watson–Crick pairing |
| $I_5$ | $j \to 5j$ | $\{1,5\},\{3,7\}$ | Wobble degeneracy |
| $I_3$ | $j \to 3j$ | $\{1,3\},\{5,7\}$ | Position-2 dominance |
The 3 six-fold amino acids (Leu, Arg, Ser) correspond 1-to-1 to the 3 involutions.
Serine uniquely crosses the purine/pyrimidine divide, requiring full charge conjugation ($I_7$) on both positions.
Reading Frame: Why 3 Bases
Bond sum: $S = 2(j_1 + \cdots + j_n) \bmod 8$
$$n = 2: S \text{ can } = 0 \text{ (dead end)} \qquad n = 3: S \in \{2,6\}, \text{ never } 0 \qquad n = 4: S \text{ can } = 0$$
$n = 3$ is the smallest odd $n$ giving sufficient combinatorial richness ($4^3 = 64$ codons).
The reading frame is a $\mathbb{Z}_8$ arithmetic inevitability.
Double Helix Uniqueness
The antiparallel double helix is the unique stable periodic
configuration on $S^2 \!\vee\! S^2$ with $\mathbb{Z}_8$ holonomy. Proved by elimination in five lemmas:
Single helix → breaks $\mathbb{Z}_2$ sphere symmetry → eliminated
$j \to 3j$ map → inconsistent winding → eliminated
Quadruple helix → $V_4$ has only 3 involutions → eliminated
Triple helix → odd strand count breaks $\mathbb{Z}_2$ → eliminated
Double helix with $I_7$ → unique survivor